Center for Biophysics and Quantitative Biology
Expanded alphabet, precise sequencing make DNA the next data storage solution

Xuenan Mi receives the Graduate Student Excellence Award from the American Chemical Society
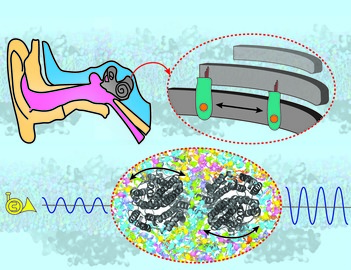
Molecular mechanism of hearing highlighted in the first atomic-resolution picture of outer hair cell surface proteins
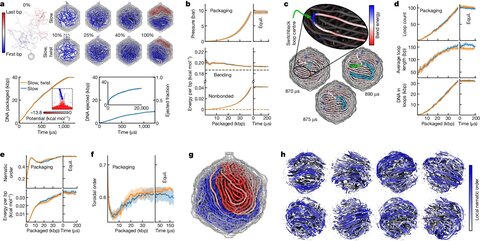
First atom-level structure of packaged viral genome reveals new properties and dynamics

Govindjee receives Lifetime Achievement Award for photosynthesis research